ChIA-DropBox: a novel analysis and visualization pipeline for multiplex chromatin interactions | bioRxiv

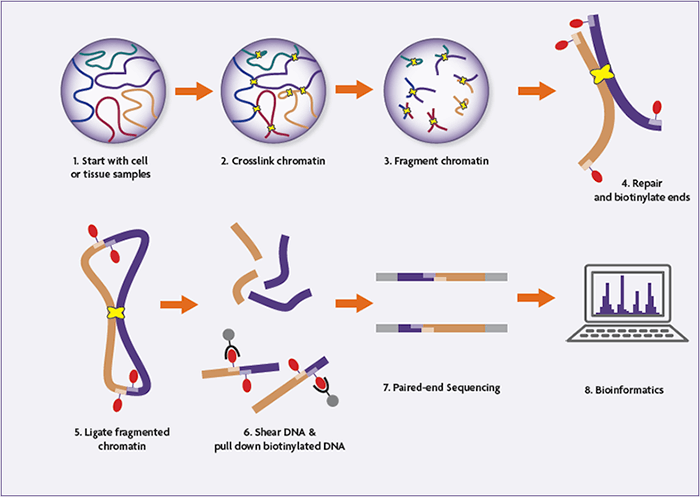
Chromatin Interaction Analysis Using Paired‐End Tag Sequencing - Fullwood - 2010 - Current Protocols in Molecular Biology - Wiley Online Library

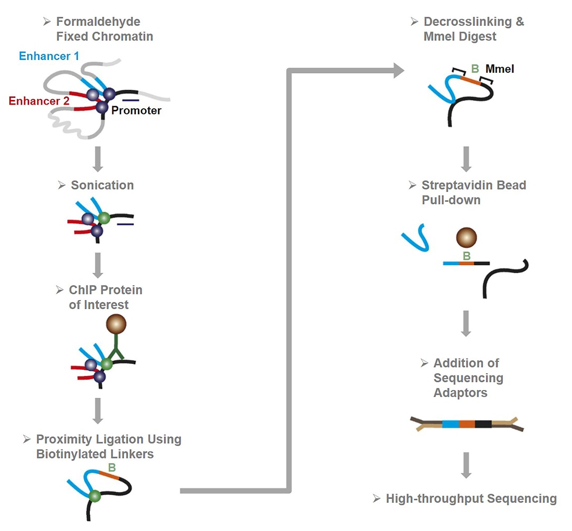
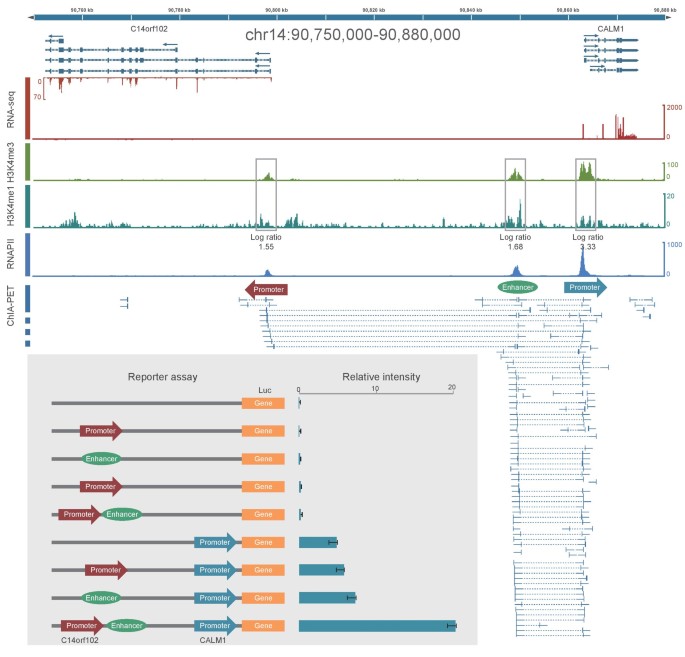
Figure 1 from Chromatin Interaction Analysis with Paired-End Tag Sequencing (ChIA-PET) for Mapping Chromatin Interactions and Understanding Transcription Regulation | Semantic Scholar
![PDF] Chromatin Interaction Analysis with Paired-End Tag Sequencing (ChIA-PET) for Mapping Chromatin Interactions and Understanding Transcription Regulation | Semantic Scholar PDF] Chromatin Interaction Analysis with Paired-End Tag Sequencing (ChIA-PET) for Mapping Chromatin Interactions and Understanding Transcription Regulation | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/002a88635940a2cf0fec7ffb53e894c93834278e/9-Figure3-1.png)
PDF] Chromatin Interaction Analysis with Paired-End Tag Sequencing (ChIA-PET) for Mapping Chromatin Interactions and Understanding Transcription Regulation | Semantic Scholar

In situ Chromatin Interaction Analysis Using Paired‐End Tag Sequencing - Wang - 2021 - Current Protocols - Wiley Online Library

ChIP-based methods for the identification of long-range chromatin interactions. - Abstract - Europe PMC
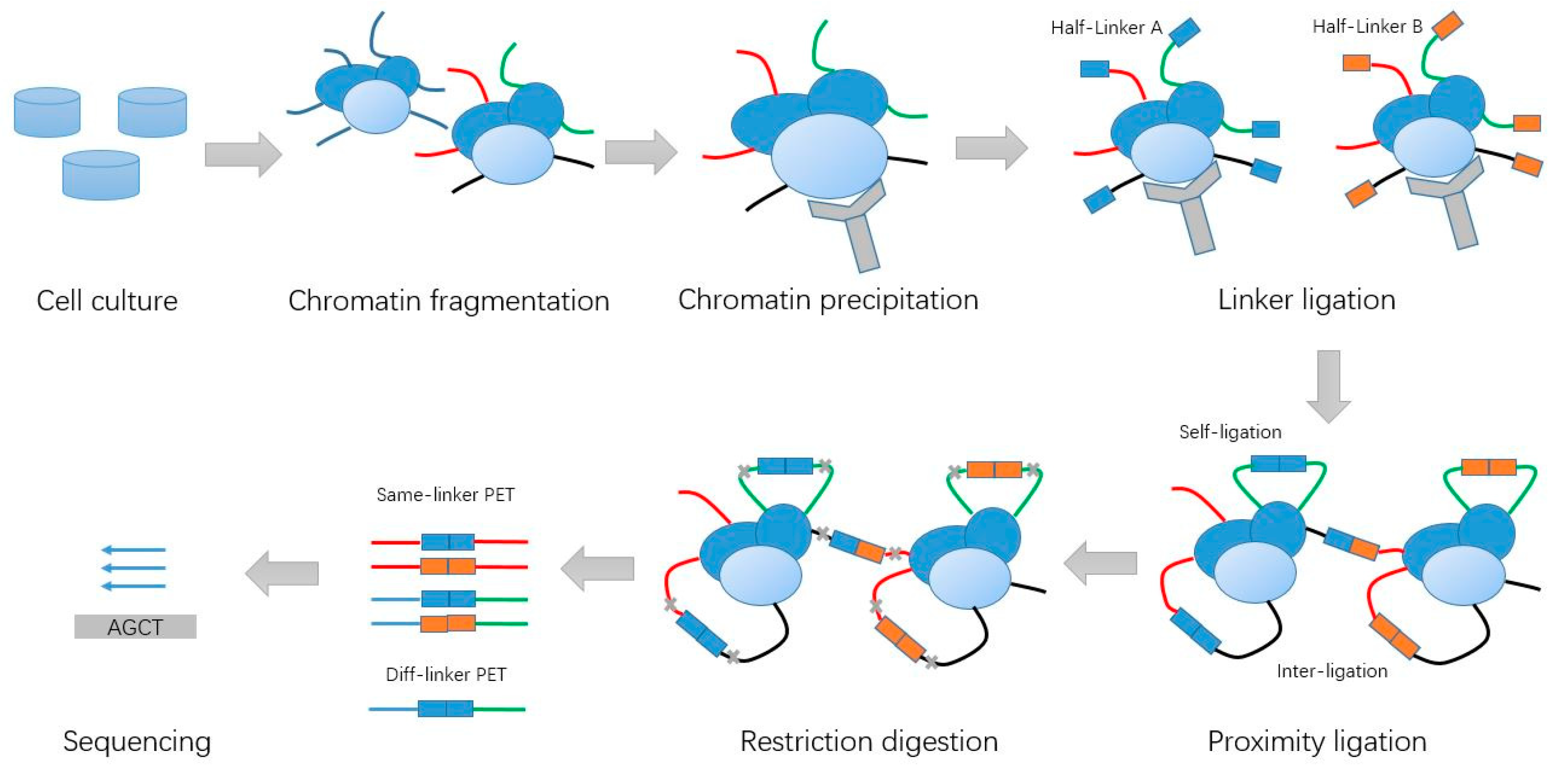
Chromatin Interaction Analysis with Paired-End Tag (ChIA-PET) sequencing technology and application | BMC Genomics | Full Text
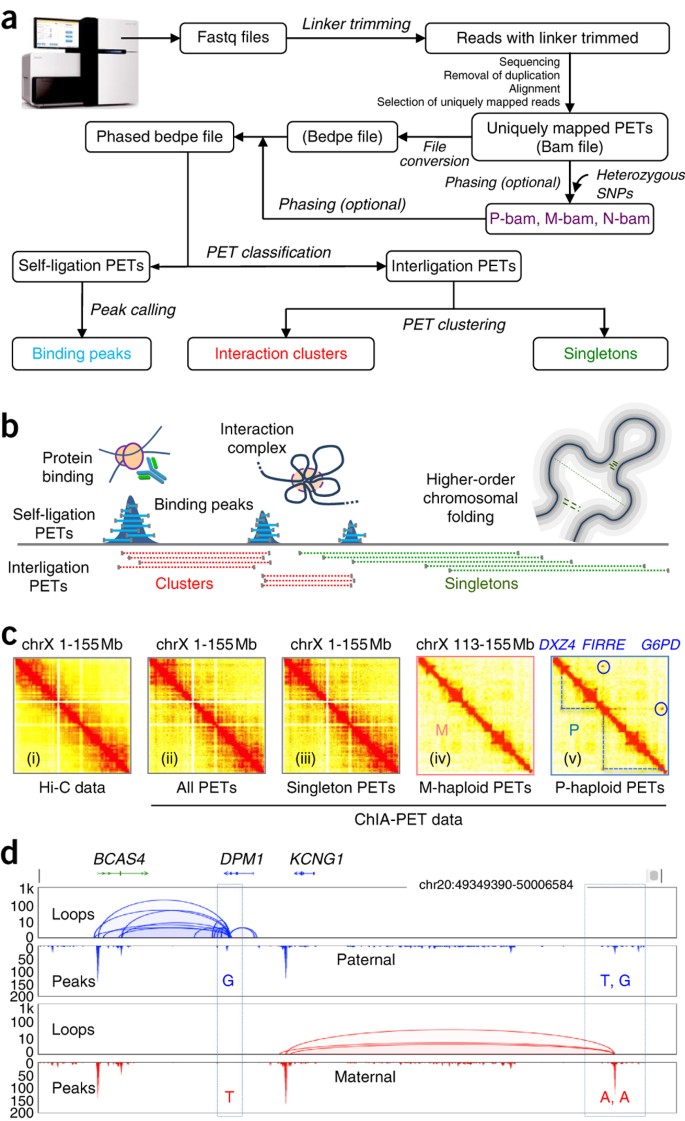
Long-read ChIA-PET for base-pair-resolution mapping of haplotype-specific chromatin interactions | Nature Protocols

Long-read ChIA-PET for base-pair-resolution mapping of haplotype-specific chromatin interactions | Nature Protocols
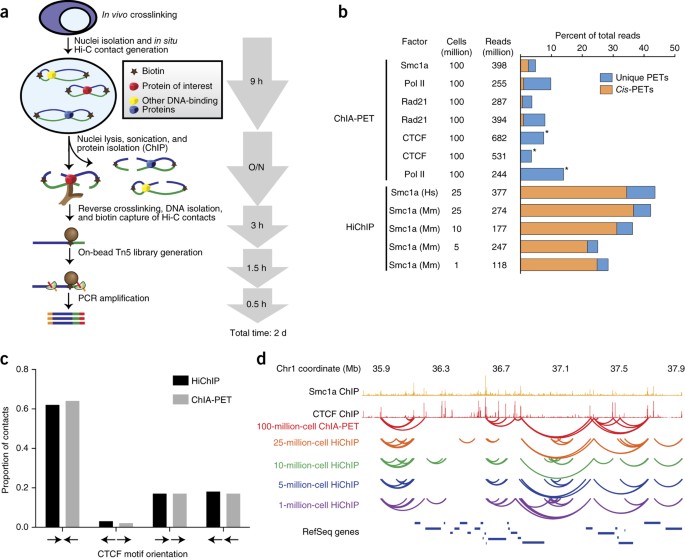
HiCuT: An efficient and low input method to identify protein-directed chromatin interactions | PLOS Genetics

Landscapes and archipelagos: spatial organization of gene regulation in vertebrates: Trends in Cell Biology

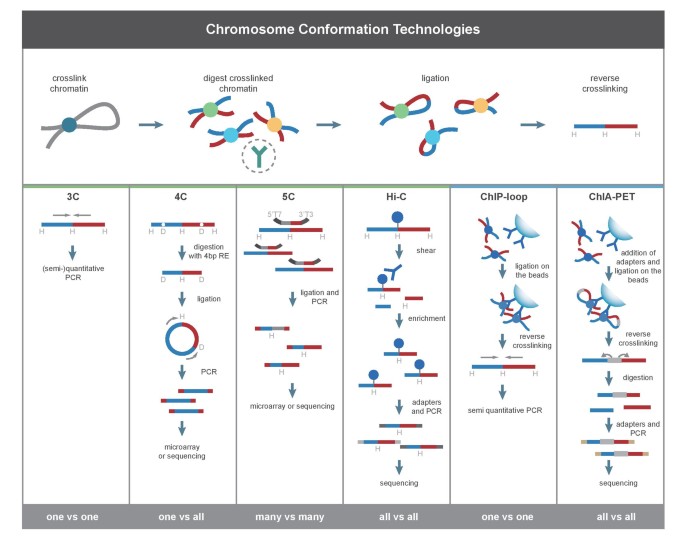
Chromosome conformation capture technologies and their impact in understanding genome function | SpringerLink